AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Chromosome scaffold protein12/26/2023  J Cell Biol 108:2435–2447īelyaev ND, Budker VG, Dubrovskaya VA, Kim AA, Kiseleva EV, Sidorov VN (1992) Localization of proteins forming the outer surface of isolated metaphase chromosomes. 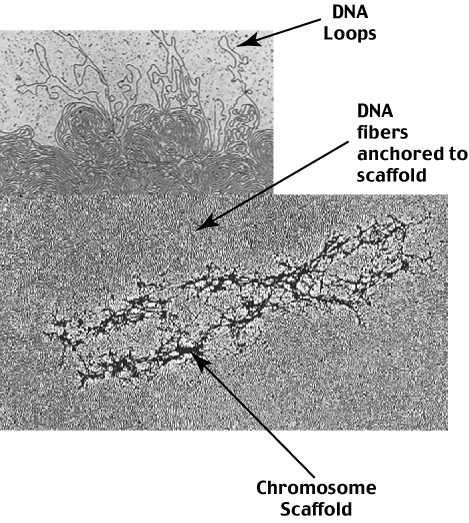
These results suggest that LFM-1 antigens fit some of the predictions of the licensing factor model, and may have a role in cell cycle dependent events.īehrens J, Mareel MM, Van-Roy FM, Birchmeier W (1989) Dissecting tumor cell invasion epithelial acquire invasive properties after the loss of uvomorulin-mediated cell-cell adhesion. They shared the sequence at the amino-terminal end but failed to show total homology with known proteins. The Mr 58000 and a minor Mr 38000 protein (which was enriched only in mitotic chromosomes of synchronized cells) were analyzed by Edman degradation. These results were independently confirmed on nuclei sorted by flow cytometry and in cell populations synchronized by release of G1-/S-phase hydroxyurea arrest. Parallel analysis of interphase nuclei revealed LFM-1 epitopes inside G1-, but excluded from G2-phase nuclei. In addition, confocal microscopy of those chromosomes revealed the LFM-1 epitopes distributed on the external surface and the axis of chromatids. Biochemical extraction procedures on isolated metaphase chromosomes from nocodazole-synchronized cells indicated that the Mr 58000 protein behaves as a chromosomal scaffold protein, that is, it remains in the pellets after high salt (2 M NaCl) or 3′–5′ diiodosalicylic acid treatments, even in DNAse pre-digested samples. Cell fractionation methods followed by immunoblotting and immunofluorescence showed that this protein is associated with the nuclear fraction. In the present work we report an Mr 58000 protein in MDCK epithelial cells, recognized by a monoclonal antibody (LFM-1) that decorates chromosomes during M-phase. We conclude that the scaffold structure is an artifact resulting from incomplete dispersion of central chromatin and aggregation of NHPs in dehistonized chromosomes.Because the mechanisms that govern mitosis are a key to the understanding of cell growth, the proteins associated with chromosomes specifically during this phase have received thorough attention. Under these circumstances, no chromosome scaffold was seen. The PTA stain showed the kinetochore plates but no central structures and (2) isolated chromosomes were partially dispersed to decrease the high concentration of chromatin in the center of the chromosome, then treated with 4 M ammonium acetate or 2 M NaCl to dehistonize them and disperse the DNA. The EDTA-regressive stain showed ribonucleoprotein particles at the periphery of the chromosomes but nothing at the center of the chromosomes. 
Two studies were done to determine if the scaffold is real or an artifact: (1) Chinese hamster mitotic cells and isolated chromosomes were examined using two protein stains -EDTA-regressive staining and phosphotungstic acid (PTA) stain. An alternative explanation is that the scaffold in whole-mount preparations represents incomplete dispersion of the high concentration of chromatin in the center of chromosomes, and when the histones are removed and the DNA dispersed, the remaining nonhistone proteins (NHPs) aggregate to form a chromosome-shaped structure. There are several troubling aspects of this concept: (1) such structures have not been seen in many previous thin-section and whole-mount electron microscopy studies of metaphase chromosomes, while they are readily seen in leptotene and zygotene chromosomes (2) such a structure poses problems for sister chromatid exchanges and (3) the published photographs show a marked variation in the amount of scaffold in different whole-mount preparations. A protein chromosome scaffold structure has been proposed that acts as a structural framework for attachment of chromosomal DNA. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
